Nickel »
PDB 3l1m-3n6n »
3mie »
Nickel in PDB 3mie: Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3)
Enzymatic activity of Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3)
All present enzymatic activity of Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3):
1.14.17.3;
1.14.17.3;
Protein crystallography data
The structure of Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3), PDB code: 3mie
was solved by
E.E.Chufan,
B.A.Eipper,
R.E.Mains,
L.M.Amzel,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 52.56 / 3.26 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 68.719, 68.991, 81.259, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.1 / 24 |
Other elements in 3mie:
The structure of Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3) also contains other interesting chemical elements:
| Copper | (Cu) | 2 atoms |
| Sodium | (Na) | 1 atom |
Nickel Binding Sites:
The binding sites of Nickel atom in the Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3)
(pdb code 3mie). This binding sites where shown within
5.0 Angstroms radius around Nickel atom.
In total only one binding site of Nickel was determined in the Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3), PDB code: 3mie:
In total only one binding site of Nickel was determined in the Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3), PDB code: 3mie:
Nickel binding site 1 out of 1 in 3mie
Go back to
Nickel binding site 1 out
of 1 in the Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3)
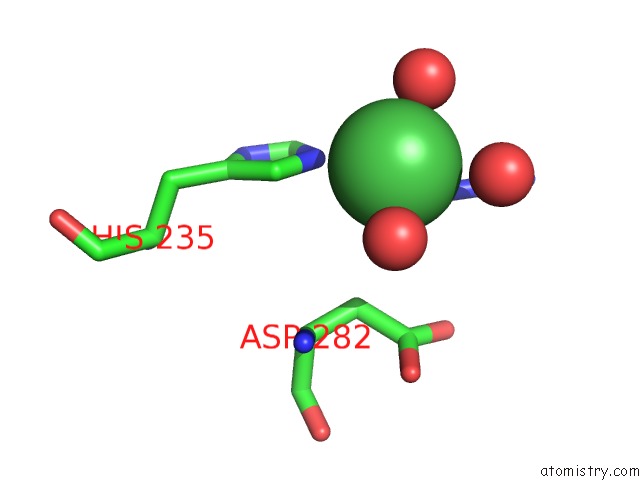
Mono view
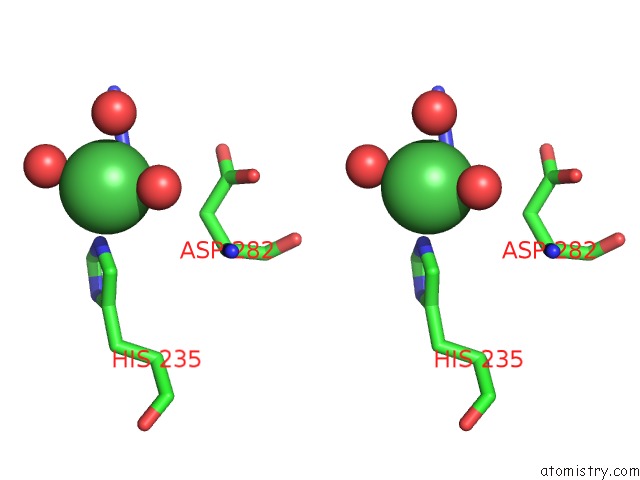
Stereo pair view

Mono view

Stereo pair view
A full contact list of Nickel with other atoms in the Ni binding
site number 1 of Oxidized (CU2+) Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) with Bound Azide Obtained By Soaking (50MM NAN3) within 5.0Å range:
|
Reference:
E.E.Chufan,
S.T.Prigge,
X.Siebert,
B.A.Eipper,
R.E.Mains,
L.M.Amzel.
Differential Reactivity Between the Two Copper Sites of Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) J.Am.Chem.Soc. V. 132 15565 2010.
ISSN: ISSN 0002-7863
PubMed: 20958070
DOI: 10.1021/JA103117R
Page generated: Mon Aug 18 18:47:12 2025
ISSN: ISSN 0002-7863
PubMed: 20958070
DOI: 10.1021/JA103117R
Last articles
Ni in 6G40Ni in 6G3Y
Ni in 6FPW
Ni in 6G3X
Ni in 6FPI
Ni in 6FPO
Ni in 6G1B
Ni in 6FET
Ni in 6FOP
Ni in 6F5S